The full dataset viewer is not available (click to read why). Only showing a preview of the rows.
Error code: DatasetGenerationCastError
Exception: DatasetGenerationCastError
Message: An error occurred while generating the dataset
All the data files must have the same columns, but at some point there are 17138 new columns ({'AP3B2', 'RP11-96L14.7', 'SMARCA4', 'NAA20', 'ZNF117', 'CCM2L', 'TBCC', 'KCNMB2-AS1', 'ACSM6', 'HAS3', 'TWF2', 'ZDHHC19', 'CACNA1C', 'YWHAE', 'CLN6', 'CCDC57', 'TAS2R5', 'PFDN2', 'SHANK2', 'GATA6', 'RACK1', 'IPMK', 'KIF5C', 'IGFBP2', 'RHOT2', 'THAP12', 'SERPINI2', 'ZNF229', 'MAPKAPK2', 'RFTN1', 'ARL9', 'GAPT', 'TAS2R20', 'FAM49B', 'PEX16', 'MYLIP', 'CYP2D6', 'ITM2C', 'ATP5J2', 'OR2A1-AS1', 'CXorf56', 'SLX1B', 'MRPL13', 'HP1BP3', 'TMEM115', 'PHB2', 'TRPV2', 'FAAP24', 'LCE6A', 'MAPK8IP2', 'SERPINB1', 'C5orf45', 'CTD-2007L18.5', 'TRIAP1', 'EXO1', 'REEP2', 'MSRB3', 'TMEM52', 'MRPL39', 'FAM156A', 'TRIM44', 'ZNF558', 'IDE', 'MATR3', 'ALDH16A1', 'SORBS3', 'ATPAF1', 'TGFBI', 'CPAMD8', 'IL15', 'CLHC1', 'CLEC12B', 'LRP11', 'SLC52A2', 'RP11-226E21.4', 'ZNF784', 'TMBIM6', 'SPERT', 'VIP', 'C2CD2', 'VGLL4', 'BMX', 'PPP1R3B', 'HNRNPH2', 'DSC2', 'BZW2', 'INPPL1', 'NPIPB11', 'RP11-679C8.2', 'ATP6V1A', 'CCDC120', 'KRT13', 'TBC1D1', 'LINGO2', 'PPP1R15B', 'UBA5', 'TMEM233', 'LRIF1', 'FAM131C', 'COIL', 'CRISPLD2', 'S100A8', 'PPEF1', 'ZNHIT3', 'TTYH1', 'FAM60A', 'H1FX', 'SGIP1', 'TM4SF18', 'ALKBH5', 'BEND6', 'PHB', 'STIM1', 'JUNB', 'IL32', 'LY6D', 'LINC01181', 'RPL41', 'TMED3', 'RGPD1', 'RP11-223P11.3', 'ATP12A', 'ALDH1A1', 'CDK5RAP1', 'TACC3', 'CIAPIN1', 'ASGR1', 'PLA2G4B', 'MREG', 'DUSP3', 'DUSP16', 'MIR1199', 'GALNT3', 'CCR5', 'SLC25A29', 'C7orf55', 'NCAPG', 'LIPE-AS1', 'SDF2', 'MSL2', 'TBC1D8', 'TREML1', 'ADHFE1', 'CHRDL1', 'CCND1', 'IL36G', 'C14orf166', 'SFTPD-AS1', 'PSMB7', 'CCS', 'KRT18',
...
', 'PNPLA5', 'NUDT19', 'SRR', 'NPTXR', 'USP20', 'ADAM15', 'COX5B', 'PSENEN', 'HOXC4', 'ATAD5', 'C1orf228', 'STRA6', 'GMIP', 'RP1-71H19.2', 'TMEM184A', 'ZNF837', 'CCDC171', 'ASUN', 'TMEM59L', 'ERGIC3', 'DNAJB5', 'SLCO3A1', 'APITD1', 'RASA4', 'KIDINS220', 'ORAI3', 'FCRL1', 'HABP4', 'LA16c-385E7.1', 'HOXD3', 'PSMD6', 'PCDHGC3', 'NXT1', 'RBM43', 'DNASE1', 'LPAR1', 'VMO1', 'ACADVL', 'POLR3A', 'DEFB106A', 'MRVI1', 'CDK6', 'QTRT1', 'AP3S2', 'EFCAB10', 'TXNDC8', 'ZC3H13', 'AGPAT3', 'CDH18', 'PSMD11', 'SON', 'PLCL2', 'OBSL1', 'IL1RL2', 'PLA2G4A', 'RP11-2C24.9', 'SYT10', 'SMAD6', 'RP11-342C20.2', 'LDHB', 'CCDC81', 'KPTN', 'GEMIN7', 'NUDCD1', 'ANKRD23', 'NEMF', 'GATSL2', 'TRIM8', 'UHRF1', 'CLIC6', 'RCOR3', 'MYCL', 'SPACA6', 'DSEL', 'TCF3', 'EGR3', 'CERCAM', 'TCTN2', 'NPEPPS', 'LINC00998', 'AHNAK2', 'ARIH2', 'EAPP', 'VWC2', 'ATP6V1E2', 'ALG9', 'ITPKC', 'FAM188B', 'STX1A', 'KALRN', 'TMEM40', 'TNFAIP8L2', 'KIF11', 'RP11-982M15.2', 'LMAN2L', 'CTC-518B2.12', 'CORO2A', 'CCDC170', 'CASP7', 'TNPO1', 'CA5B', 'GALNT14', 'PROCA1', 'PCDHB4', 'PUSL1', 'PLAC1', 'ZNF680', 'DOCK1', 'ZDHHC15', 'LRRC39', 'RXFP1', 'AMOTL1', 'GOLGA6L4', 'RRBP1', 'KDM4C', 'CHMP6', 'RP11-368I7.4', 'ALPK3', 'IRAK4', 'PEBP1', 'ACTR2', 'RPRD1B', 'RP3-402G11.26', 'UNC5A', 'CYB561D1', 'AC104024.1', 'LIMS3', 'TLE2', 'TRIM26', 'KIAA1551', 'AIG1', 'DUSP8', 'PPP2R2D', 'USP36', 'KCNIP4', 'MID2', 'HIST1H1E', 'ARHGAP40', 'NAA15', 'DDX58', 'CAPN1', 'CAMK2B', 'FAM188A', 'SIN3B', 'TMED5', 'PRR26', 'OGFOD1', 'BEX3', 'CAPG', 'TPCN1', 'RPA2'}) and 3 missing columns ({'yaxis', 'xaxis', 'r'}).
This happened while the csv dataset builder was generating data using
hf://datasets/jiawennnn/STimage-1K4M/ST/gene_exp/GSE144239_GSM4284316_count.csv (at revision 1c2a89874eed9bb70606373d2b7551c9583a0131)
Please either edit the data files to have matching columns, or separate them into different configurations (see docs at https://hf.co/docs/hub/datasets-manual-configuration#multiple-configurations)
Traceback: Traceback (most recent call last):
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 1870, in _prepare_split_single
writer.write_table(table)
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/arrow_writer.py", line 622, in write_table
pa_table = table_cast(pa_table, self._schema)
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/table.py", line 2292, in table_cast
return cast_table_to_schema(table, schema)
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/table.py", line 2240, in cast_table_to_schema
raise CastError(
datasets.table.CastError: Couldn't cast
Unnamed: 0: string
MIR1302-2: double
RP11-34P13.7: double
RP11-34P13.14: double
FO538757.1: double
RP4-669L17.10: double
RP11-206L10.9: double
LINC00115: double
RP11-54O7.1: double
SAMD11: double
NOC2L: double
KLHL17: double
PLEKHN1: double
PERM1: double
HES4: double
ISG15: double
AGRN: double
RNF223: double
C1orf159: double
TTLL10: double
TNFRSF18: double
TNFRSF4: double
SDF4: double
B3GALT6: double
FAM132A: double
UBE2J2: double
SCNN1D: double
ACAP3: double
PUSL1: double
CPSF3L: double
CPTP: double
TAS1R3: double
DVL1: double
MXRA8: double
AURKAIP1: double
CCNL2: double
MRPL20: double
RP4-758J18.13: double
ANKRD65: double
VWA1: double
ATAD3C: double
ATAD3B: double
ATAD3A: double
TMEM240: double
SSU72: double
FNDC10: double
MIB2: double
MMP23B: double
CDK11B: double
SLC35E2B: double
CDK11A: double
SLC35E2: double
NADK: double
GNB1: double
RP1-140A9.1: double
CALML6: double
TMEM52: double
CFAP74: double
GABRD: double
PRKCZ: double
FAAP20: double
SKI: double
MORN1: double
RER1: double
PEX10: double
PLCH2: double
PANK4: double
HES5: double
TNFRSF14: double
MMEL1: double
FAM213B: double
TTC34: double
PRDM16: double
ARHGEF16: double
MEGF6: double
RP11-46F15.2: double
TPRG1L: double
WRAP73: double
TP73: double
SMIM1: double
LRRC47: double
CEP104: double
DFFB: double
C1orf174: double
AJAP1: double
NPHP4: double
KCNAB2: double
RP1-120G22.11: double
RPL22: double
RNF207: double
ICMT: double
GPR153: double
ACOT7: double
HES2: double
ESPN: double
TNFRSF25: double
PLEKHG5: double
NOL9:
...
e
NSDHL: double
ZNF185: double
PNMA3: double
PNMA6A: double
LL0XNC01-16G2.1: double
ZNF275: double
ZFP92: double
TREX2: double
HAUS7: double
BGN: double
FAM58A: double
DUSP9: double
SLC6A8: double
BCAP31: double
ABCD1: double
SRPK3: double
IDH3G: double
SSR4: double
PDZD4: double
AVPR2: double
L1CAM: double
ARHGAP4: double
NAA10: double
RENBP: double
HCFC1: double
TMEM187: double
IRAK1: double
MECP2: double
TKTL1: double
FLNA: double
EMD: double
DNASE1L1: double
RPL10: double
TAZ: double
ATP6AP1: double
GDI1: double
FAM50A: double
PLXNA3: double
LAGE3: double
UBL4A: double
SLC10A3: double
FAM3A: double
G6PD: double
IKBKG: double
CTAG1A: double
CTAG1B: double
CTAG2: double
GAB3: double
DKC1: double
MPP1: double
H2AFB1: double
F8A1: double
F8: double
FUNDC2: double
BRCC3: double
MTCP1: double
VBP1: double
CLIC2: double
H2AFB2: double
F8A2: double
F8A3: double
H2AFB3: double
TMLHE: double
SPRY3: double
VAMP7: double
IL9R: double
RPS4Y1: double
ZFY: double
LINC00278: double
PCDH11Y: double
TBL1Y: double
USP9Y: double
DDX3Y: double
UTY: double
TMSB4Y: double
NLGN4Y: double
FAM224B: double
FAM224A: double
HSFY1: double
HSFY2: double
TTTY14: double
KDM5D: double
EIF1AY: double
RPS4Y2: double
BPY2: double
DAZ1: double
DAZ2: double
AC012005.2: double
AC012005.1: double
BPY2B: double
DAZ3: double
DAZ4: double
BPY2C: double
AC006386.1: double
LINC00266-4P: double
AC006328.1: double
-- schema metadata --
pandas: '{"index_columns": [{"kind": "range", "name": null, "start": 0, "' + 1929671
to
{'Unnamed: 0': Value(dtype='string', id=None), 'yaxis': Value(dtype='float64', id=None), 'xaxis': Value(dtype='float64', id=None), 'r': Value(dtype='float64', id=None)}
because column names don't match
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "/src/services/worker/src/worker/job_runners/config/parquet_and_info.py", line 1415, in compute_config_parquet_and_info_response
parquet_operations, partial, estimated_dataset_info = stream_convert_to_parquet(
File "/src/services/worker/src/worker/job_runners/config/parquet_and_info.py", line 991, in stream_convert_to_parquet
builder._prepare_split(
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 1741, in _prepare_split
for job_id, done, content in self._prepare_split_single(
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 1872, in _prepare_split_single
raise DatasetGenerationCastError.from_cast_error(
datasets.exceptions.DatasetGenerationCastError: An error occurred while generating the dataset
All the data files must have the same columns, but at some point there are 17138 new columns ({'AP3B2', 'RP11-96L14.7', 'SMARCA4', 'NAA20', 'ZNF117', 'CCM2L', 'TBCC', 'KCNMB2-AS1', 'ACSM6', 'HAS3', 'TWF2', 'ZDHHC19', 'CACNA1C', 'YWHAE', 'CLN6', 'CCDC57', 'TAS2R5', 'PFDN2', 'SHANK2', 'GATA6', 'RACK1', 'IPMK', 'KIF5C', 'IGFBP2', 'RHOT2', 'THAP12', 'SERPINI2', 'ZNF229', 'MAPKAPK2', 'RFTN1', 'ARL9', 'GAPT', 'TAS2R20', 'FAM49B', 'PEX16', 'MYLIP', 'CYP2D6', 'ITM2C', 'ATP5J2', 'OR2A1-AS1', 'CXorf56', 'SLX1B', 'MRPL13', 'HP1BP3', 'TMEM115', 'PHB2', 'TRPV2', 'FAAP24', 'LCE6A', 'MAPK8IP2', 'SERPINB1', 'C5orf45', 'CTD-2007L18.5', 'TRIAP1', 'EXO1', 'REEP2', 'MSRB3', 'TMEM52', 'MRPL39', 'FAM156A', 'TRIM44', 'ZNF558', 'IDE', 'MATR3', 'ALDH16A1', 'SORBS3', 'ATPAF1', 'TGFBI', 'CPAMD8', 'IL15', 'CLHC1', 'CLEC12B', 'LRP11', 'SLC52A2', 'RP11-226E21.4', 'ZNF784', 'TMBIM6', 'SPERT', 'VIP', 'C2CD2', 'VGLL4', 'BMX', 'PPP1R3B', 'HNRNPH2', 'DSC2', 'BZW2', 'INPPL1', 'NPIPB11', 'RP11-679C8.2', 'ATP6V1A', 'CCDC120', 'KRT13', 'TBC1D1', 'LINGO2', 'PPP1R15B', 'UBA5', 'TMEM233', 'LRIF1', 'FAM131C', 'COIL', 'CRISPLD2', 'S100A8', 'PPEF1', 'ZNHIT3', 'TTYH1', 'FAM60A', 'H1FX', 'SGIP1', 'TM4SF18', 'ALKBH5', 'BEND6', 'PHB', 'STIM1', 'JUNB', 'IL32', 'LY6D', 'LINC01181', 'RPL41', 'TMED3', 'RGPD1', 'RP11-223P11.3', 'ATP12A', 'ALDH1A1', 'CDK5RAP1', 'TACC3', 'CIAPIN1', 'ASGR1', 'PLA2G4B', 'MREG', 'DUSP3', 'DUSP16', 'MIR1199', 'GALNT3', 'CCR5', 'SLC25A29', 'C7orf55', 'NCAPG', 'LIPE-AS1', 'SDF2', 'MSL2', 'TBC1D8', 'TREML1', 'ADHFE1', 'CHRDL1', 'CCND1', 'IL36G', 'C14orf166', 'SFTPD-AS1', 'PSMB7', 'CCS', 'KRT18',
...
', 'PNPLA5', 'NUDT19', 'SRR', 'NPTXR', 'USP20', 'ADAM15', 'COX5B', 'PSENEN', 'HOXC4', 'ATAD5', 'C1orf228', 'STRA6', 'GMIP', 'RP1-71H19.2', 'TMEM184A', 'ZNF837', 'CCDC171', 'ASUN', 'TMEM59L', 'ERGIC3', 'DNAJB5', 'SLCO3A1', 'APITD1', 'RASA4', 'KIDINS220', 'ORAI3', 'FCRL1', 'HABP4', 'LA16c-385E7.1', 'HOXD3', 'PSMD6', 'PCDHGC3', 'NXT1', 'RBM43', 'DNASE1', 'LPAR1', 'VMO1', 'ACADVL', 'POLR3A', 'DEFB106A', 'MRVI1', 'CDK6', 'QTRT1', 'AP3S2', 'EFCAB10', 'TXNDC8', 'ZC3H13', 'AGPAT3', 'CDH18', 'PSMD11', 'SON', 'PLCL2', 'OBSL1', 'IL1RL2', 'PLA2G4A', 'RP11-2C24.9', 'SYT10', 'SMAD6', 'RP11-342C20.2', 'LDHB', 'CCDC81', 'KPTN', 'GEMIN7', 'NUDCD1', 'ANKRD23', 'NEMF', 'GATSL2', 'TRIM8', 'UHRF1', 'CLIC6', 'RCOR3', 'MYCL', 'SPACA6', 'DSEL', 'TCF3', 'EGR3', 'CERCAM', 'TCTN2', 'NPEPPS', 'LINC00998', 'AHNAK2', 'ARIH2', 'EAPP', 'VWC2', 'ATP6V1E2', 'ALG9', 'ITPKC', 'FAM188B', 'STX1A', 'KALRN', 'TMEM40', 'TNFAIP8L2', 'KIF11', 'RP11-982M15.2', 'LMAN2L', 'CTC-518B2.12', 'CORO2A', 'CCDC170', 'CASP7', 'TNPO1', 'CA5B', 'GALNT14', 'PROCA1', 'PCDHB4', 'PUSL1', 'PLAC1', 'ZNF680', 'DOCK1', 'ZDHHC15', 'LRRC39', 'RXFP1', 'AMOTL1', 'GOLGA6L4', 'RRBP1', 'KDM4C', 'CHMP6', 'RP11-368I7.4', 'ALPK3', 'IRAK4', 'PEBP1', 'ACTR2', 'RPRD1B', 'RP3-402G11.26', 'UNC5A', 'CYB561D1', 'AC104024.1', 'LIMS3', 'TLE2', 'TRIM26', 'KIAA1551', 'AIG1', 'DUSP8', 'PPP2R2D', 'USP36', 'KCNIP4', 'MID2', 'HIST1H1E', 'ARHGAP40', 'NAA15', 'DDX58', 'CAPN1', 'CAMK2B', 'FAM188A', 'SIN3B', 'TMED5', 'PRR26', 'OGFOD1', 'BEX3', 'CAPG', 'TPCN1', 'RPA2'}) and 3 missing columns ({'yaxis', 'xaxis', 'r'}).
This happened while the csv dataset builder was generating data using
hf://datasets/jiawennnn/STimage-1K4M/ST/gene_exp/GSE144239_GSM4284316_count.csv (at revision 1c2a89874eed9bb70606373d2b7551c9583a0131)
Please either edit the data files to have matching columns, or separate them into different configurations (see docs at https://hf.co/docs/hub/datasets-manual-configuration#multiple-configurations)Need help to make the dataset viewer work? Make sure to review how to configure the dataset viewer, and open a discussion for direct support.
Unnamed: 0
string | yaxis
float64 | xaxis
float64 | r
float64 |
|---|---|---|---|
GSE144239_GSM4284316_10x26 | 5,741.5 | 3,845.3 | 45.125581 |
GSE144239_GSM4284316_10x28 | 6,142.9 | 3,841.4 | 45.125581 |
GSE144239_GSM4284316_10x30 | 6,549.5 | 3,834.4 | 45.125581 |
GSE144239_GSM4284316_10x32 | 6,951.6 | 3,839.6 | 45.125581 |
GSE144239_GSM4284316_10x34 | 7,355.5 | 3,839.4 | 45.125581 |
GSE144239_GSM4284316_10x36 | 7,756.1 | 3,839.3 | 45.125581 |
GSE144239_GSM4284316_10x38 | 8,144.4 | 3,838.8 | 45.125581 |
GSE144239_GSM4284316_10x40 | 8,554.1 | 3,833.4 | 45.125581 |
GSE144239_GSM4284316_10x44 | 9,356.4 | 3,830.9 | 45.125581 |
GSE144239_GSM4284316_10x46 | 9,758 | 3,828.8 | 45.125581 |
GSE144239_GSM4284316_10x48 | 10,156 | 3,830.9 | 45.125581 |
GSE144239_GSM4284316_11x25 | 5,551.5 | 4,022.2 | 45.125581 |
GSE144239_GSM4284316_11x27 | 5,940.9 | 4,020.2 | 45.125581 |
GSE144239_GSM4284316_11x29 | 6,353 | 4,009.4 | 45.125581 |
GSE144239_GSM4284316_11x31 | 6,757.6 | 4,014.4 | 45.125581 |
GSE144239_GSM4284316_11x33 | 7,155.4 | 4,013.1 | 45.125581 |
GSE144239_GSM4284316_11x35 | 7,557.4 | 4,014.1 | 45.125581 |
GSE144239_GSM4284316_11x37 | 7,947.4 | 4,004.7 | 45.125581 |
GSE144239_GSM4284316_11x39 | 8,337.8 | 4,010.5 | 45.125581 |
GSE144239_GSM4284316_11x41 | 8,760.5 | 4,011.8 | 45.125581 |
GSE144239_GSM4284316_11x43 | 9,164.1 | 4,015.6 | 45.125581 |
GSE144239_GSM4284316_11x45 | 9,564 | 4,006.9 | 45.125581 |
GSE144239_GSM4284316_11x47 | 9,959.5 | 4,018.4 | 45.125581 |
GSE144239_GSM4284316_11x49 | 10,355.8 | 4,014.1 | 45.125581 |
GSE144239_GSM4284316_12x22 | 4,940.1 | 4,194.9 | 45.125581 |
GSE144239_GSM4284316_12x24 | 5,340.2 | 4,194.8 | 45.125581 |
GSE144239_GSM4284316_12x26 | 5,747.8 | 4,194.1 | 45.125581 |
GSE144239_GSM4284316_12x28 | 6,143.3 | 4,195.9 | 45.125581 |
GSE144239_GSM4284316_12x30 | 6,554.3 | 4,190.9 | 45.125581 |
GSE144239_GSM4284316_12x32 | 6,951.6 | 4,195.8 | 45.125581 |
GSE144239_GSM4284316_12x34 | 7,354.3 | 4,197.5 | 45.125581 |
GSE144239_GSM4284316_12x36 | 7,750.3 | 4,200.9 | 45.125581 |
GSE144239_GSM4284316_12x38 | 8,142.6 | 4,186.9 | 45.125581 |
GSE144239_GSM4284316_12x40 | 8,553.2 | 4,187.2 | 45.125581 |
GSE144239_GSM4284316_12x42 | 8,954.5 | 4,190.6 | 45.125581 |
GSE144239_GSM4284316_12x44 | 9,357.6 | 4,191.9 | 45.125581 |
GSE144239_GSM4284316_12x46 | 9,757.8 | 4,194.4 | 45.125581 |
GSE144239_GSM4284316_12x48 | 10,154.5 | 4,193.3 | 45.125581 |
GSE144239_GSM4284316_13x21 | 4,734.1 | 4,378 | 45.125581 |
GSE144239_GSM4284316_13x23 | 5,143 | 4,373.2 | 45.125581 |
GSE144239_GSM4284316_13x25 | 5,539.8 | 4,382.1 | 45.125581 |
GSE144239_GSM4284316_13x27 | 5,933.8 | 4,384.9 | 45.125581 |
GSE144239_GSM4284316_13x29 | 6,348.2 | 4,374.1 | 45.125581 |
GSE144239_GSM4284316_13x31 | 6,745.5 | 4,377.5 | 45.125581 |
GSE144239_GSM4284316_13x33 | 7,148.5 | 4,380.3 | 45.125581 |
GSE144239_GSM4284316_13x35 | 7,555.3 | 4,379 | 45.125581 |
GSE144239_GSM4284316_13x37 | 7,949.6 | 4,375.2 | 45.125581 |
GSE144239_GSM4284316_13x39 | 8,347.3 | 4,372.9 | 45.125581 |
GSE144239_GSM4284316_13x41 | 8,752.6 | 4,376.6 | 45.125581 |
GSE144239_GSM4284316_13x43 | 9,160.5 | 4,375.2 | 45.125581 |
GSE144239_GSM4284316_13x45 | 9,558.7 | 4,368.6 | 45.125581 |
GSE144239_GSM4284316_13x47 | 9,955.3 | 4,374.2 | 45.125581 |
GSE144239_GSM4284316_13x49 | 10,358.7 | 4,382.5 | 45.125581 |
GSE144239_GSM4284316_14x18 | 4,130.1 | 4,567 | 45.125581 |
GSE144239_GSM4284316_14x20 | 4,522.4 | 4,558.1 | 45.125581 |
GSE144239_GSM4284316_14x22 | 4,925.8 | 4,555.8 | 45.125581 |
GSE144239_GSM4284316_14x24 | 5,326.6 | 4,555.5 | 45.125581 |
GSE144239_GSM4284316_14x26 | 5,727 | 4,560.2 | 45.125581 |
GSE144239_GSM4284316_14x28 | 6,132.8 | 4,554.1 | 45.125581 |
GSE144239_GSM4284316_14x30 | 6,524.2 | 4,550.7 | 45.125581 |
GSE144239_GSM4284316_14x32 | 6,928.3 | 4,554.7 | 45.125581 |
GSE144239_GSM4284316_14x34 | 7,329 | 4,559.4 | 45.125581 |
GSE144239_GSM4284316_14x36 | 7,729.1 | 4,558.3 | 45.125581 |
GSE144239_GSM4284316_14x38 | 8,128.4 | 4,552.6 | 45.125581 |
GSE144239_GSM4284316_14x40 | 8,527.1 | 4,555.9 | 45.125581 |
GSE144239_GSM4284316_14x42 | 8,929.7 | 4,557.5 | 45.125581 |
GSE144239_GSM4284316_14x44 | 9,328.8 | 4,559.7 | 45.125581 |
GSE144239_GSM4284316_14x46 | 9,731.5 | 4,549.9 | 45.125581 |
GSE144239_GSM4284316_14x48 | 10,129.5 | 4,554.9 | 45.125581 |
GSE144239_GSM4284316_14x50 | 10,531.1 | 4,556.3 | 45.125581 |
GSE144239_GSM4284316_15x17 | 3,939.2 | 4,743.3 | 45.125581 |
GSE144239_GSM4284316_15x19 | 4,327.9 | 4,745.6 | 45.125581 |
GSE144239_GSM4284316_15x21 | 4,726.3 | 4,734.8 | 45.125581 |
GSE144239_GSM4284316_15x23 | 5,128.5 | 4,735.4 | 45.125581 |
GSE144239_GSM4284316_15x25 | 5,536.5 | 4,736.6 | 45.125581 |
GSE144239_GSM4284316_15x27 | 5,931.5 | 4,741.7 | 45.125581 |
GSE144239_GSM4284316_15x29 | 6,329.8 | 4,730 | 45.125581 |
GSE144239_GSM4284316_15x31 | 6,729.5 | 4,734.1 | 45.125581 |
GSE144239_GSM4284316_15x33 | 7,130.9 | 4,738.7 | 45.125581 |
GSE144239_GSM4284316_15x35 | 7,532.7 | 4,739.5 | 45.125581 |
GSE144239_GSM4284316_15x37 | 7,931.4 | 4,729.9 | 45.125581 |
GSE144239_GSM4284316_15x39 | 8,322 | 4,738.1 | 45.125581 |
GSE144239_GSM4284316_15x41 | 8,731 | 4,736.8 | 45.125581 |
GSE144239_GSM4284316_15x43 | 9,131.5 | 4,740 | 45.125581 |
GSE144239_GSM4284316_15x45 | 9,533.9 | 4,733.1 | 45.125581 |
GSE144239_GSM4284316_15x47 | 9,933.6 | 4,733.6 | 45.125581 |
GSE144239_GSM4284316_15x49 | 10,329.9 | 4,735.9 | 45.125581 |
GSE144239_GSM4284316_15x51 | 10,734.1 | 4,738.6 | 45.125581 |
GSE144239_GSM4284316_16x16 | 3,732.4 | 4,919.1 | 45.125581 |
GSE144239_GSM4284316_16x18 | 4,132.2 | 4,926.6 | 45.125581 |
GSE144239_GSM4284316_16x20 | 4,527.4 | 4,922.8 | 45.125581 |
GSE144239_GSM4284316_16x22 | 4,925.8 | 4,917.7 | 45.125581 |
GSE144239_GSM4284316_16x24 | 5,333.2 | 4,915.9 | 45.125581 |
GSE144239_GSM4284316_16x26 | 5,724.5 | 4,925.1 | 45.125581 |
GSE144239_GSM4284316_16x28 | 6,132.7 | 4,914 | 45.125581 |
GSE144239_GSM4284316_16x30 | 6,527.9 | 4,915.3 | 45.125581 |
GSE144239_GSM4284316_16x32 | 6,930.5 | 4,918.4 | 45.125581 |
GSE144239_GSM4284316_16x34 | 7,330 | 4,917.3 | 45.125581 |
GSE144239_GSM4284316_16x36 | 7,733 | 4,916.8 | 45.125581 |
GSE144239_GSM4284316_16x38 | 8,127.2 | 4,913 | 45.125581 |
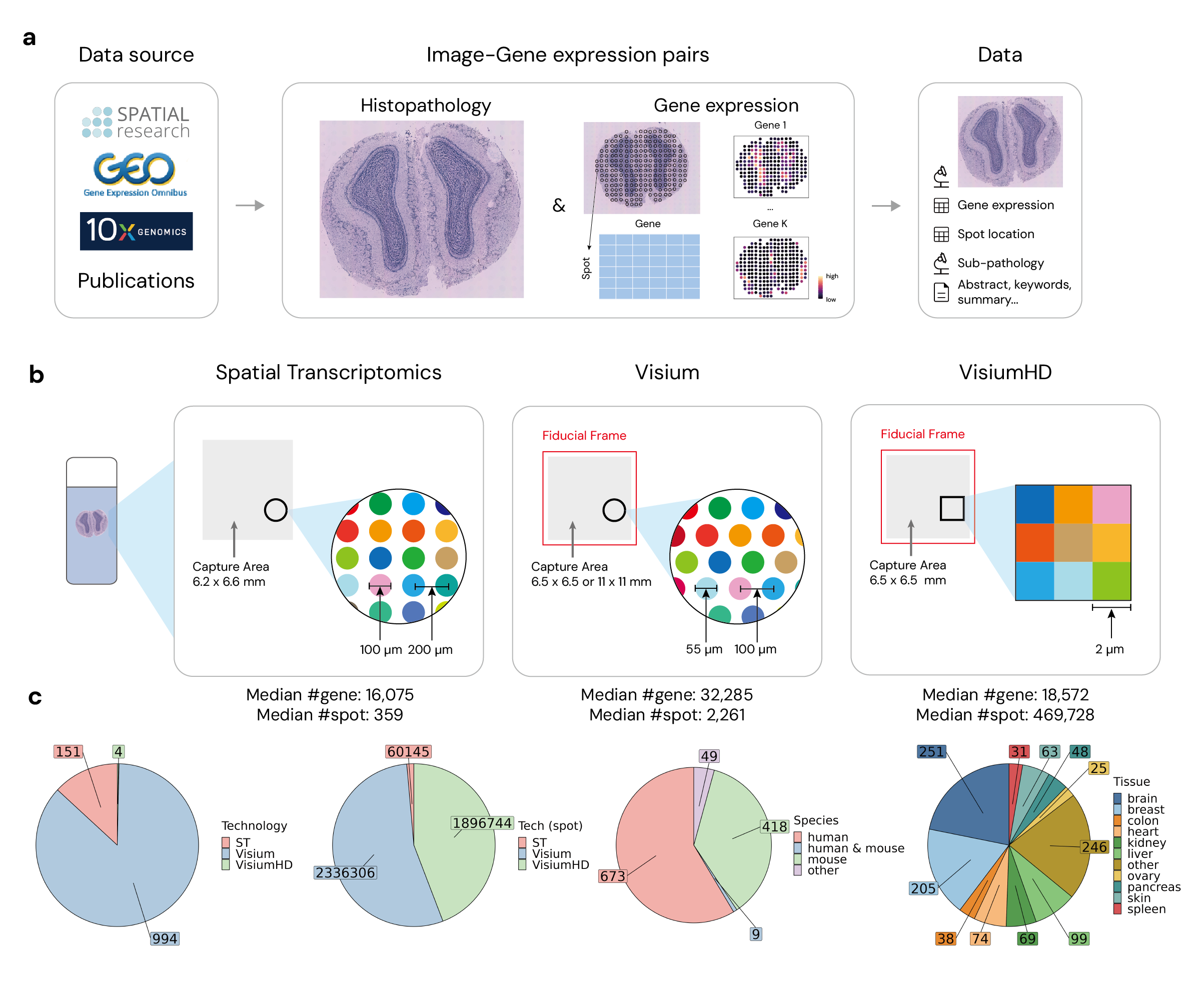
STimage-1K4M Dataset
Welcome to the STimage-1K4M Dataset repository. This dataset is designed to foster research in the field of spatial transcriptomics, combining high-resolution histopathology images with detailed gene expression data.
Update
Feb 12, 2025 We corrected a typo in meta file (changed "Human_Brain+Kidney_10X_02212023_Visium" to "Mouse_Brain+Kidney_10X_02212023_Visium"). Please refer to meta_all_gene02122025.csv for the newest meta data.
Dataset Description
STimage-1K4M consists of 1,149 spatial transcriptomics slides, totaling over 4 million spots with paired gene expression data. This dataset includes:
- Images.
- Gene expression profiles matched with high-resolution histopathology images.
- Spatial coordinates for each spot.
Data structure
The data structure is organized as follows:
├── annotation # Pathologist annotation
├── meta # Test files (alternatively `spec` or `tests`)
│ ├── bib.txt # the bibtex for all studies with pmid included in the dataset
│ ├── meta_all_gene.csv # The meta information
├── ST # Include all data for tech: Spatial Transcriptomics
│ ├── coord # Include the spot coordinates & spot radius of each slide
│ ├── gene_exp # Include the gene expression of each slide
│ └── image # Include the image each slide
├── Visium # Include all data for tech: Visium, same structure as ST
├── VisiumHD # Include all data for tech: VisiumHD, same structure as ST
Repository structure
The code for data processing and reproducing evaluation result in the paper are in Document.
Acknowledgement
The fine-tuning and evaluation codes borrows heavily from CLIP and PLIP.
Citation
@misc{chen2024stimage1k4m,
title={STimage-1K4M: A histopathology image-gene expression dataset for spatial transcriptomics},
author={Jiawen Chen and Muqing Zhou and Wenrong Wu and Jinwei Zhang and Yun Li and Didong Li},
year={2024},
eprint={2406.06393},
archivePrefix={arXiv},
primaryClass={cs.CV}
}
License
All code is licensed under the MIT License - see the LICENSE.md file for details.
- Downloads last month
- 32,541